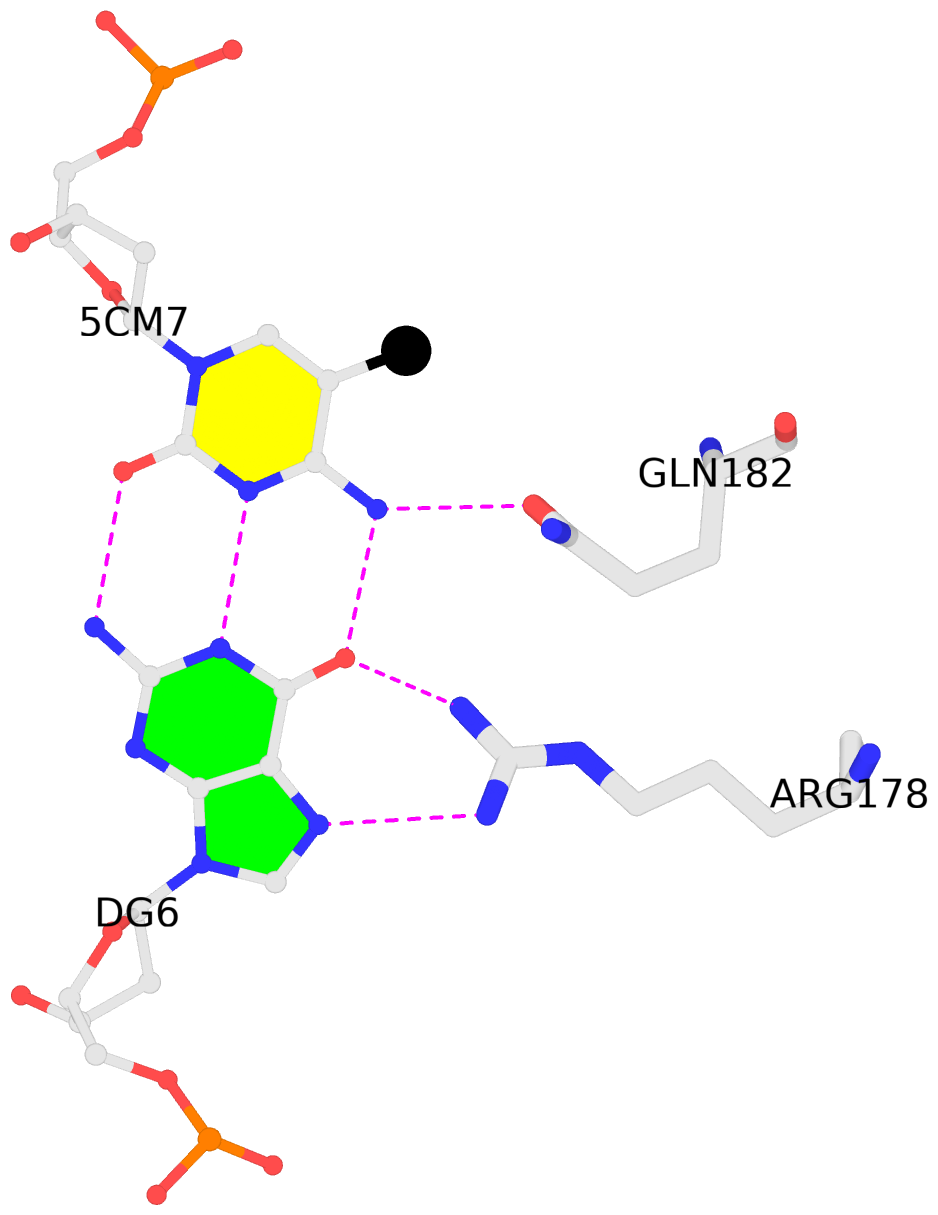
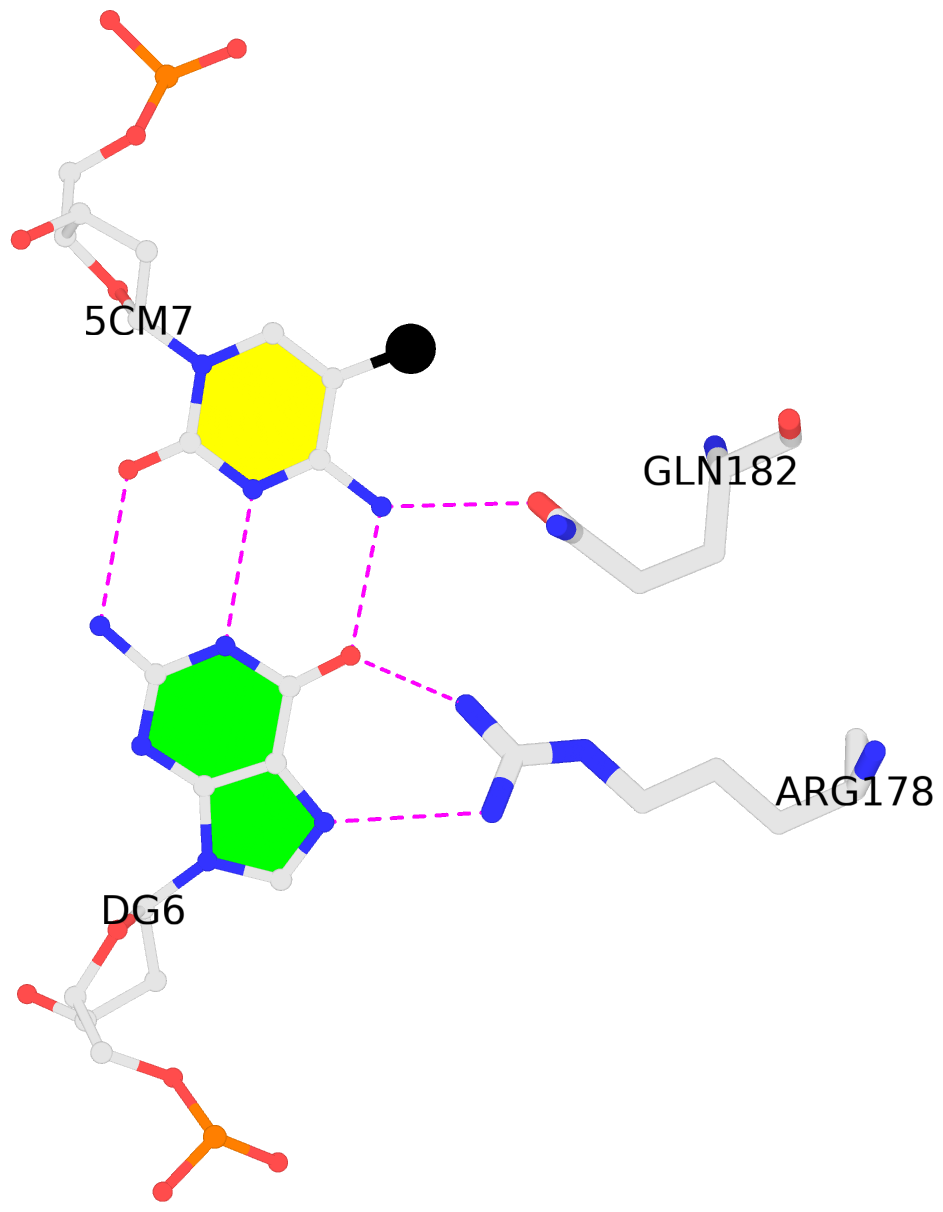
5mC interactions in PDB entry 4M9V auto-curated with SNAP
Last updated on 2019-09-30 by Xiang-Jun Lu <xiangjun@x3dna.org>. The block schematics were created with DSSR and rendered using PyMOL.
Summary information and primary citation [schematics · contacts · top · homepage · tutorial]
- PDB-id
- 4M9V
- Class
- transcription-DNA
- Method
- X-ray (0.97 Å)
- Summary
- Zfp57 mutant (e182q) in complex with 5-carboxylcytosine DNA
List of 2 5mC-amino acid contacts:-
A.5CM7: other-contacts is-WC-paired is-in-duplex [+]:CcG/cGG
-
D.5CM7: other-contacts is-WC-paired is-in-duplex [+]:CcG/cGG
-
- Reference
- Liu, Y., Olanrewaju, Y.O., Zhang, X., Cheng, X.: (2013) "DNA recognition of 5-carboxylcytosine by a zfp57 mutant at an atomic resolution of 0.97 angstrom." Biochemistry, 52, 9310-9317.
- Abstract
- The Zfp57 gene encodes a KRAB (Krüppel-associated box) domain-containing C2H2 zinc finger transcription factor that is expressed in early development. Zfp57 protein recognizes methylated CpG dinucleotide within GCGGCA elements at multiple imprinting control regions. In the previously determined structure of the mouse Zfp57 DNA-binding domain in complex with DNA containing 5-methylcytosine (5mC), the side chains of Arg178 and Glu182 contact the methyl group via hydrophobic and van der Waals interactions. We examined the role of Glu182 in recognition of 5mC by mutagenesis. The majority of mutants examined lose selectivity of methylated (5mC) over unmodified (C) and oxidative derivatives, 5-hydroxymethylcytosine, 5-formylcytosine, and 5-carboxylcytosine (5caC), suggesting that the side chain of Glu182 (the size and the charge) is dispensable for methyl group recognition but negatively impacts the binding of unmodified cytosine as well as oxidized derivatives of 5mC to achieve 5mC selectivity. Substitution of Glu182 with its corresponding amide (E182Q) had no effect on methylated DNA binding but gained significant binding affinity for 5caC DNA, resulting in a binding affinity for 5caC DNA comparable to that of the wild-type protein for 5mC. We show structurally that the uncharged amide group of E182Q interacts favorably with the carboxylate group of 5caC. Furthermore, introducing a positively charged arginine at position 182 resulted in a mutant (E182R) having higher selectivity for the negatively charged 5caC.
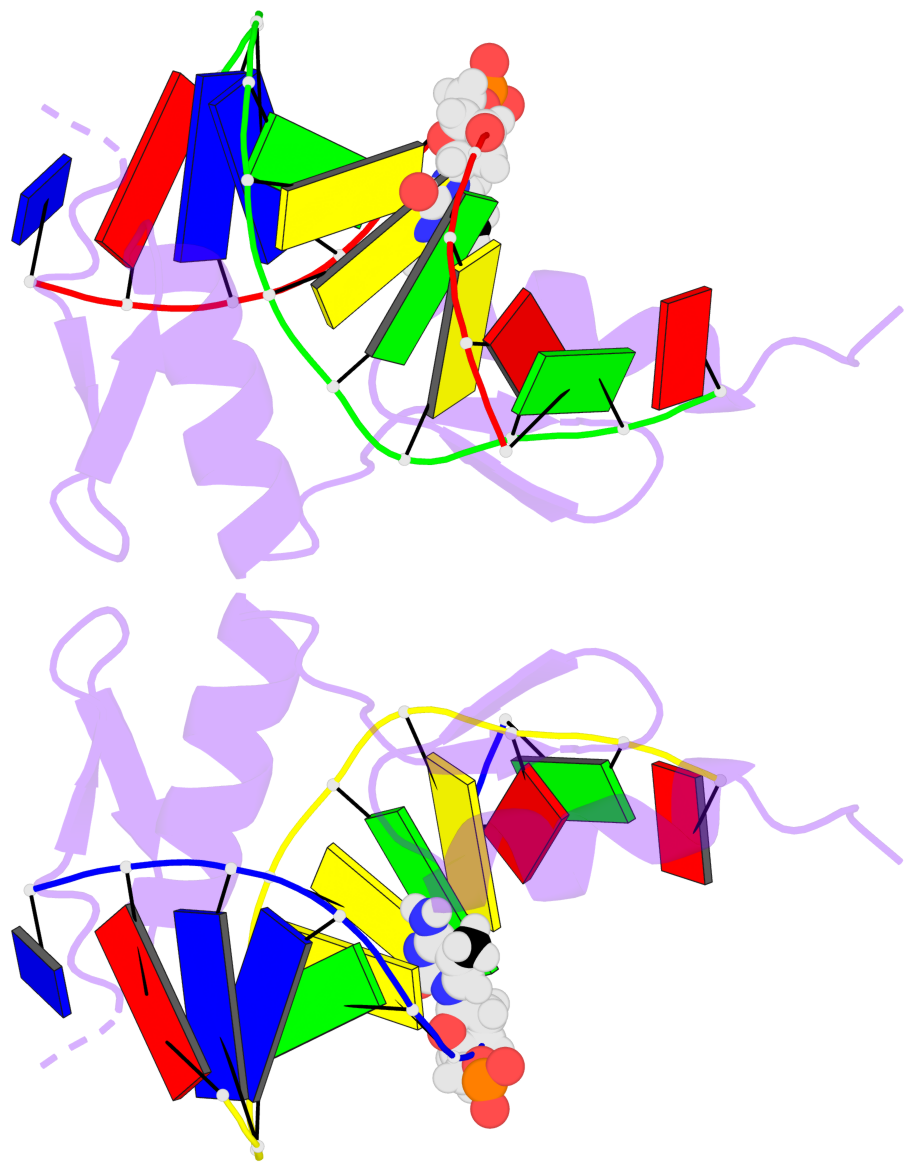
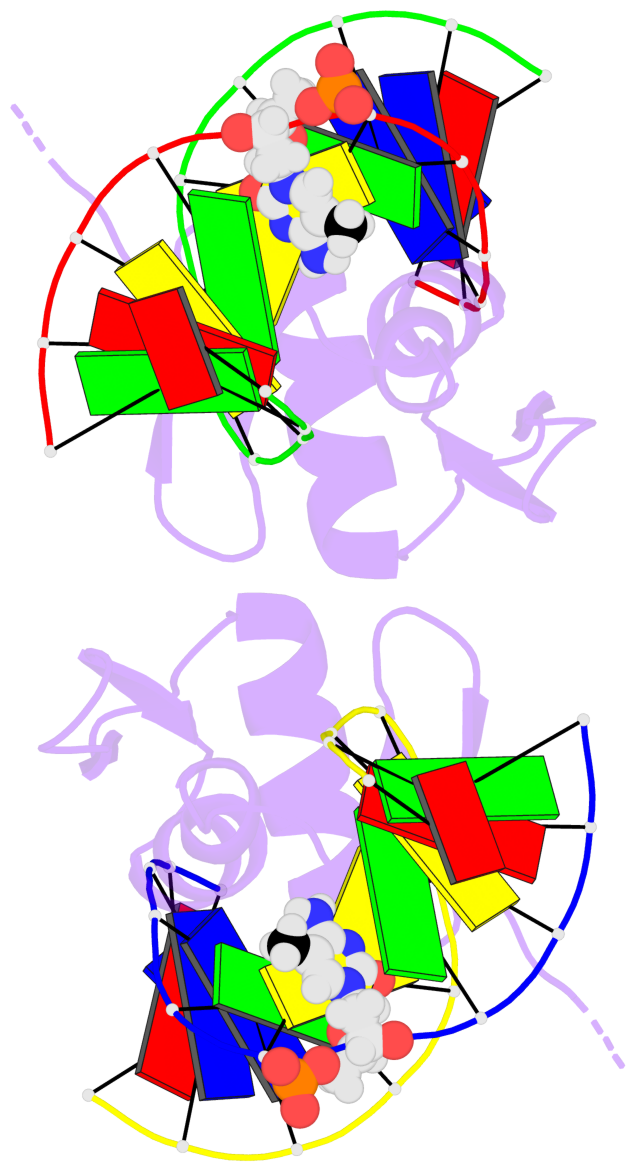
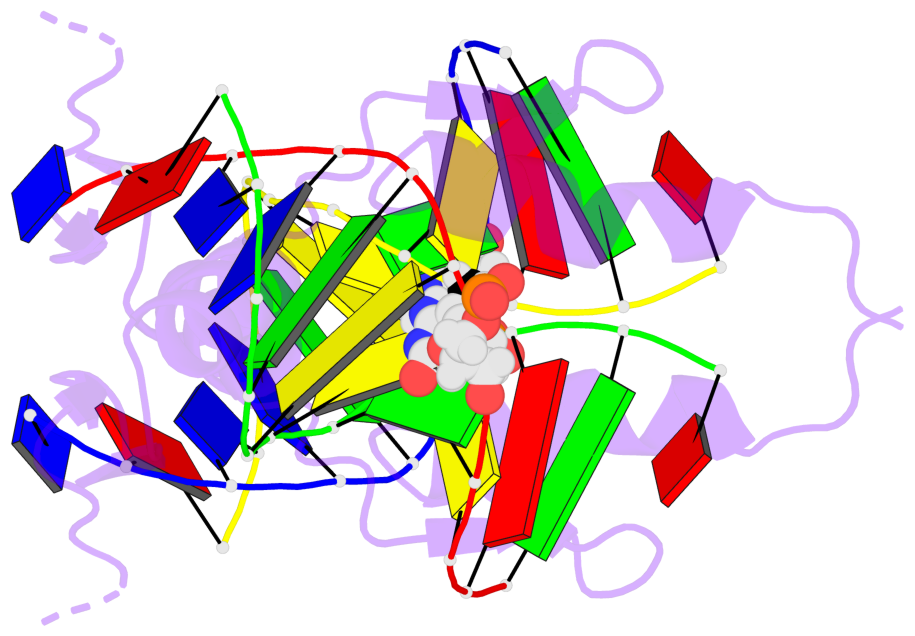
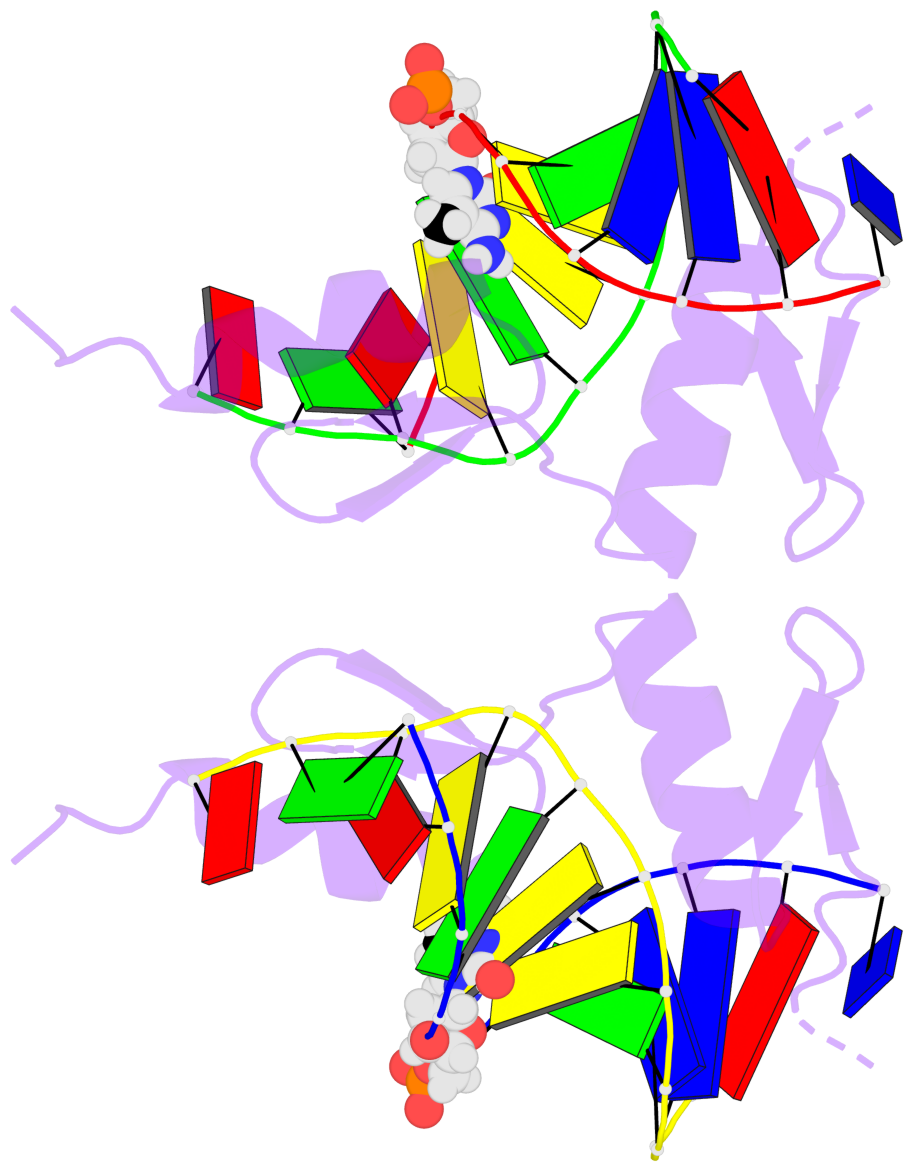
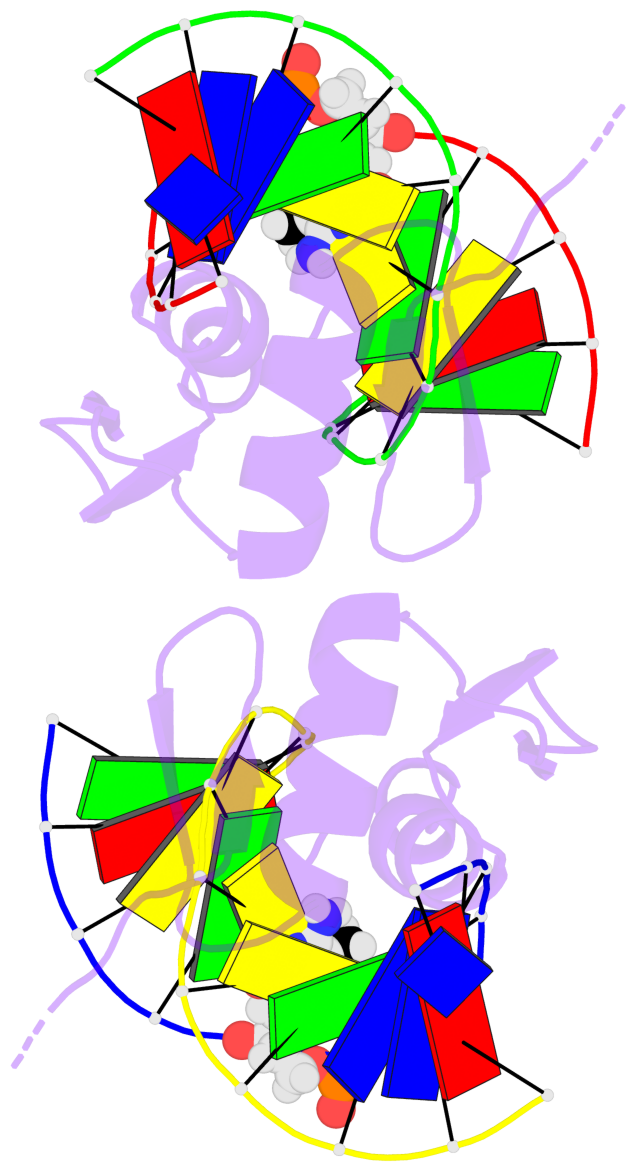
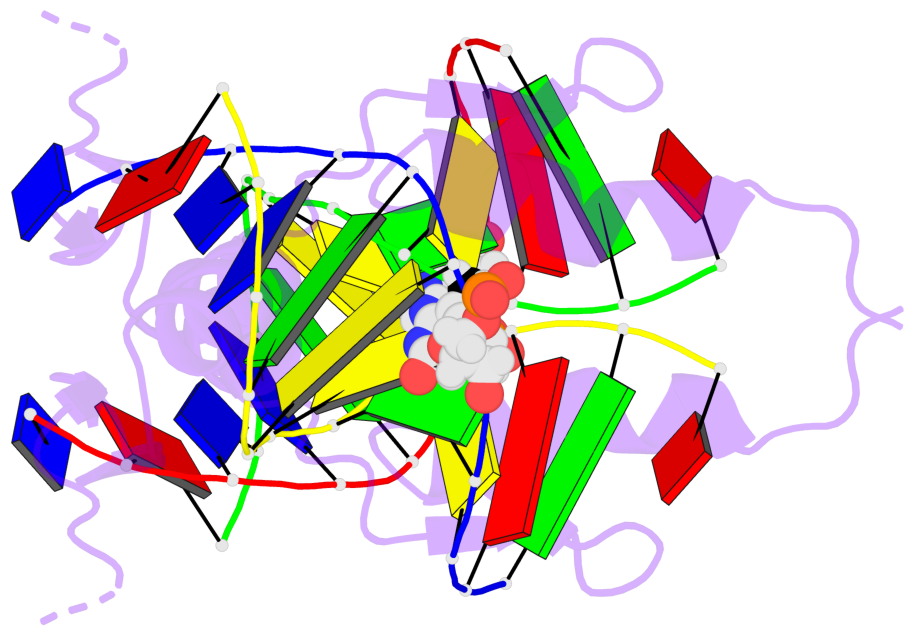
Base-block schematics in six views [summary · contacts · top · homepage · tutorial]
- The 5-methylcytosine group (PDB ligand '5CM') is shown in space-filling model, with the methyl-carbon atom in black.
- Watson-Crick base pairs are represented as long rectangular blocks with the minor-groove edge in black. Color code: A-T red, C-G yellow, G-C green, T-A blue.
- Protein is shown as cartoon in purple. DNA backbones are shown ribbon, colored code by chain identifier.
- The block schematics were created with 3DNA-DSSR, and images were rendered using PyMOL.
- Download the PyMOL session file corresponding to the top-left image in the following panel.
 |
 |
 |
 |
 |
 |
List of 2 5mC-amino acid contacts [summary · schematics · top · homepage · tutorial]
- The contacts include paired nucleotides (mostly a G in G-C pairing), and amino-acids within a 4.5-A distance cutoff to the base atoms of 5mC.
- The structure is oriented in the 'standard' base reference frame of 5mC, allowing for easy comparison and direct superimposition between entries.
- The black sphere (•) denotes the 5-methyl carbon atom in 5mC.